Anisotropic Parameters in Liquid State NMR
A New Level of Structure Determination
Maybe the most exciting application of NMR spectroscopy is structure determination in solution. Classical structural parameters are chemical shifts, J-couplings and distances derived from nuclear Overhauser enhancement (NOE) measured in isotropic liquids. However, much more structural information can be obtained if solute molecules are dissolved in anisotropic liquid environments, like liquid crystalline phases or stretched gels.
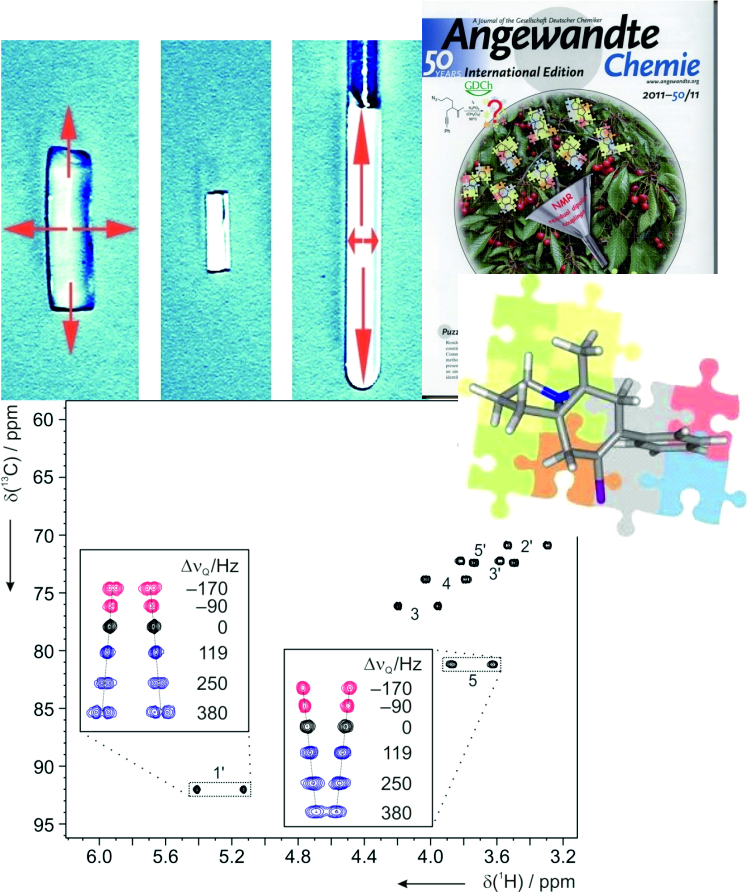
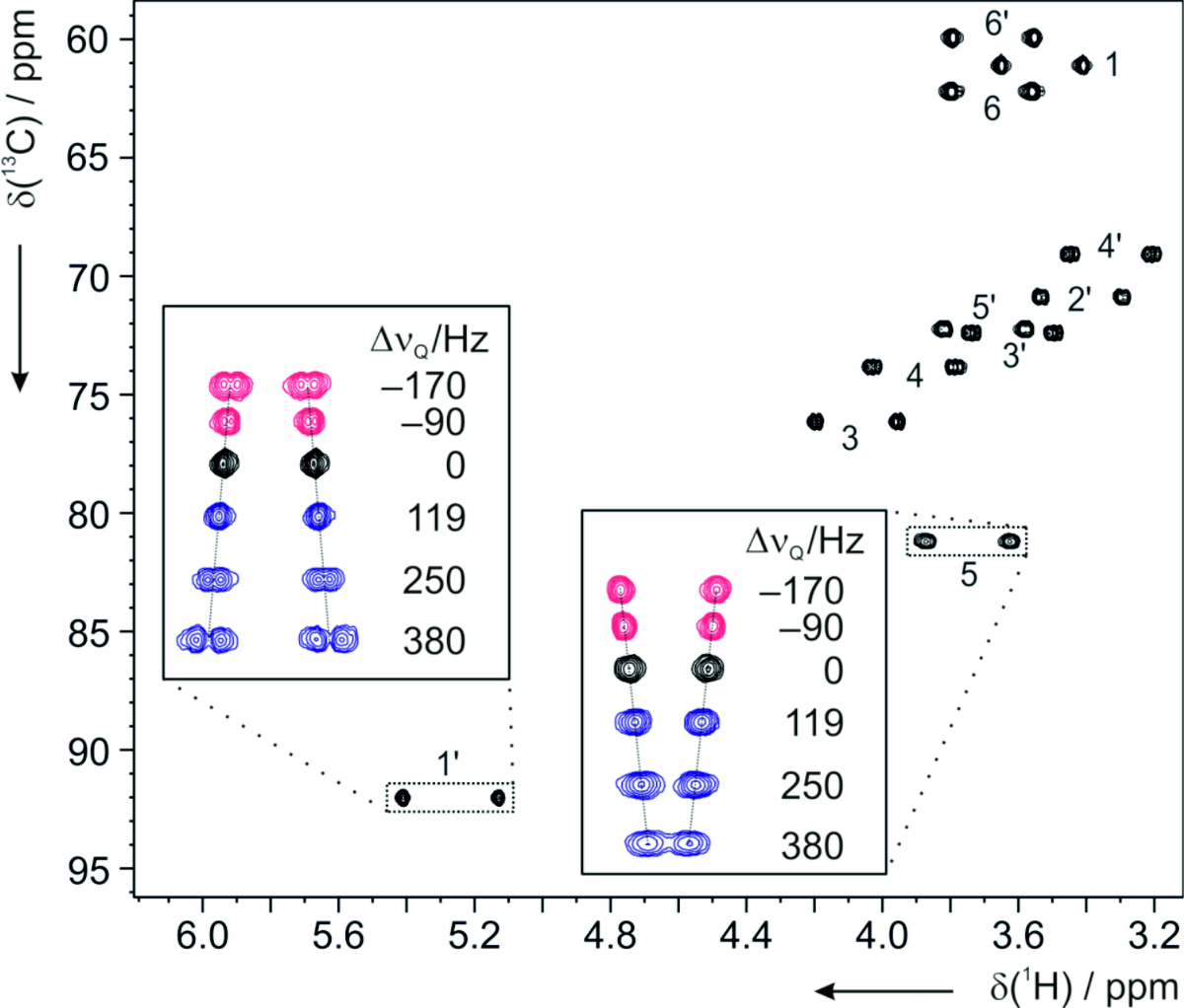
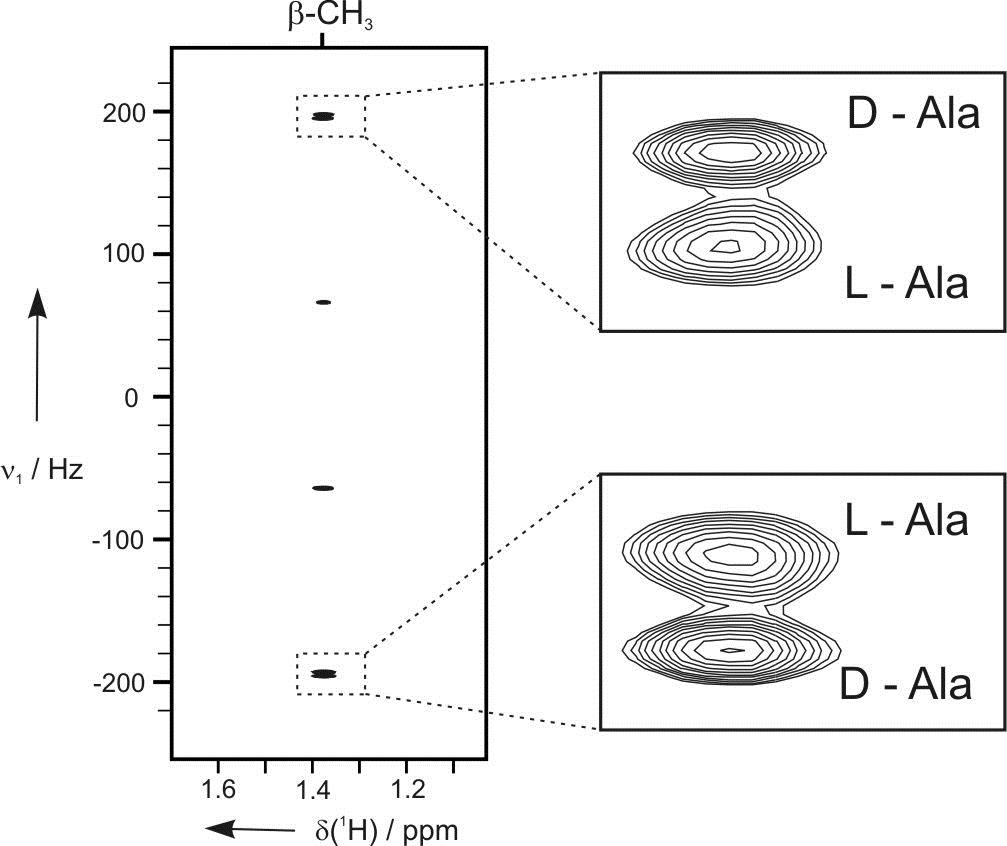
The free molecular tumbling in solution effectively averages all anisotropic parameters to zero. If partial alignment is introduced, full averaging cannot take place and some residual anisotropic parameters can be detected. The most widely used are the so-called Residual Dipolar Couplings (RDCs). They depend on the distance between interacting nuclei and on the angle between internuclear vector and external magnetic field. Thus, RDCs contain highly valuable structural information. Other residual anisotropic NMR parameters are Residual Chemical Shift Anisotropies (RCSAs) and Residual Quadrupolar Couplings (RQCs). We aim to provide methods to make anisotropic NMR parameters amenable to structure elucidation comprising constitution, configuration, conformation, and dynamics. As such we develop new ways to effectively align molecules, design pulse sequences to measure new classes of anisotropic parameters, and explore novel software tools for structure determination.
RDCs
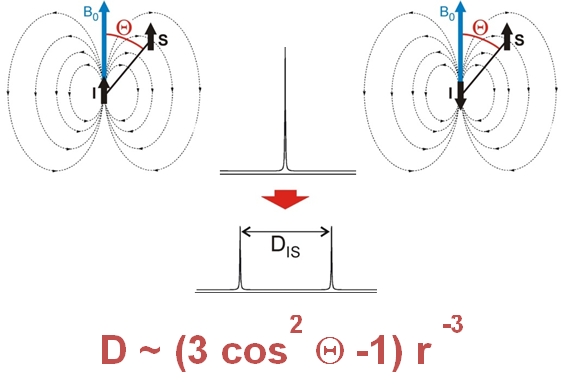
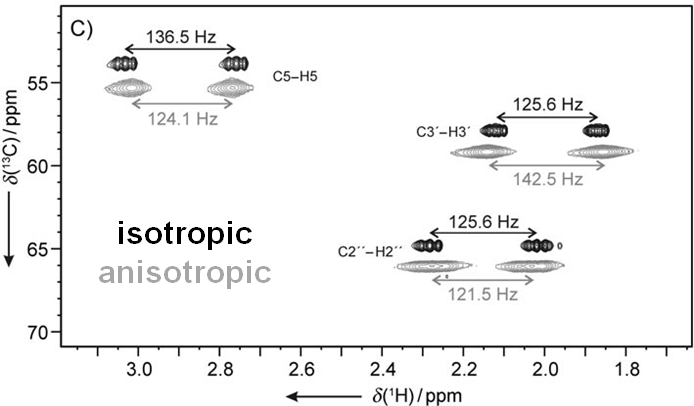
Residual dipolar couplings depend on the internuclear distance between the two NMR active nuclei (r) and the angle of the internuclear vector with respect to the outer magnetic field. Knowledge of RDCs allows the determination of the molecule’s alignment relative to an external reference, thus correlating also distant parts of the molecule. The RDCs have been used to study the conformation, configuration, constitution, and enantiomeric distinction of small-to-medium-sized organic molecules.
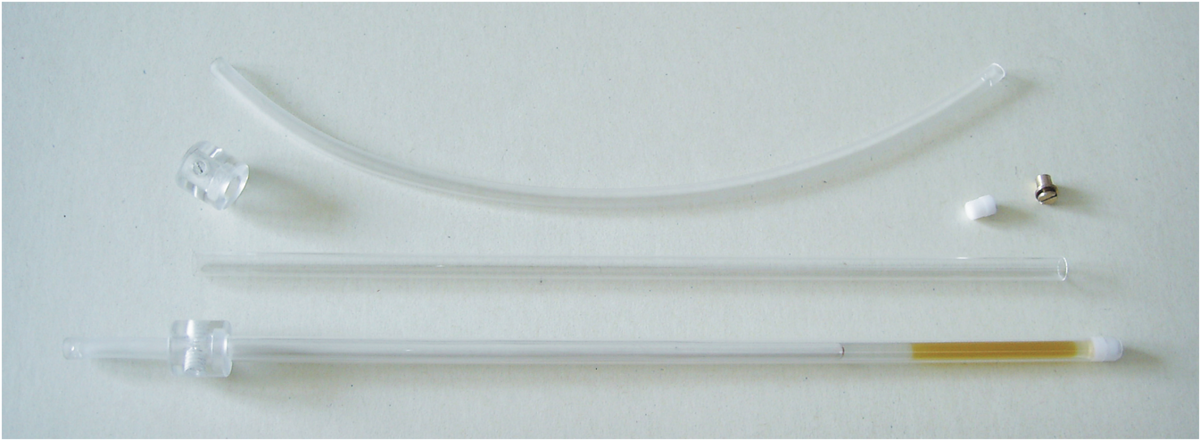
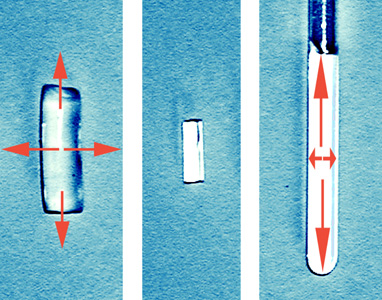
The successful measurement of RDCs requires suitable alignment media which allow proper scaling and solvents range. For organic molecules usually two types of media are used: liquid crystalline phases and stretched polymer gels. A number of polymer gel based alignment media for various solvents has been developed in our group and currently we are working on further optimisation of the alignment media.
The experimental RDCs are further used to solve different structural problems. During RDC analysis, an alignment tensor is calculated for a sterically fixed orientational model. Agreement between experimental and back calculated RDCs allows to solve the constitution, to assign the configuration, and the conformation of the molecule. The majority of the organic molecules we have recently been working with, contains some certain degree of flexibility. In such cases, the straightforward approach, which assumes a single rigid conformation, fails. Therefore, we have started to evaluate our experimental RDCs with a specialized molecular dynamics approach (see more in the Computational chemistry).