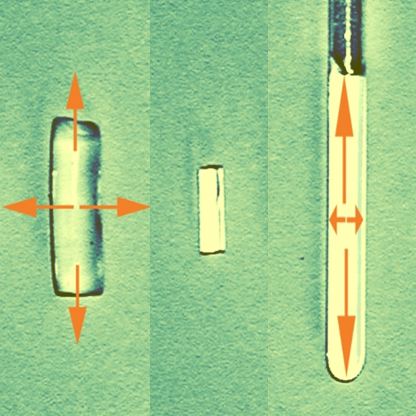

Anisotropic NMR Spectroscopy
Enabling measurement of residual anisotropic NMR parameters for the determination of configuration, conformation, and constitution.
link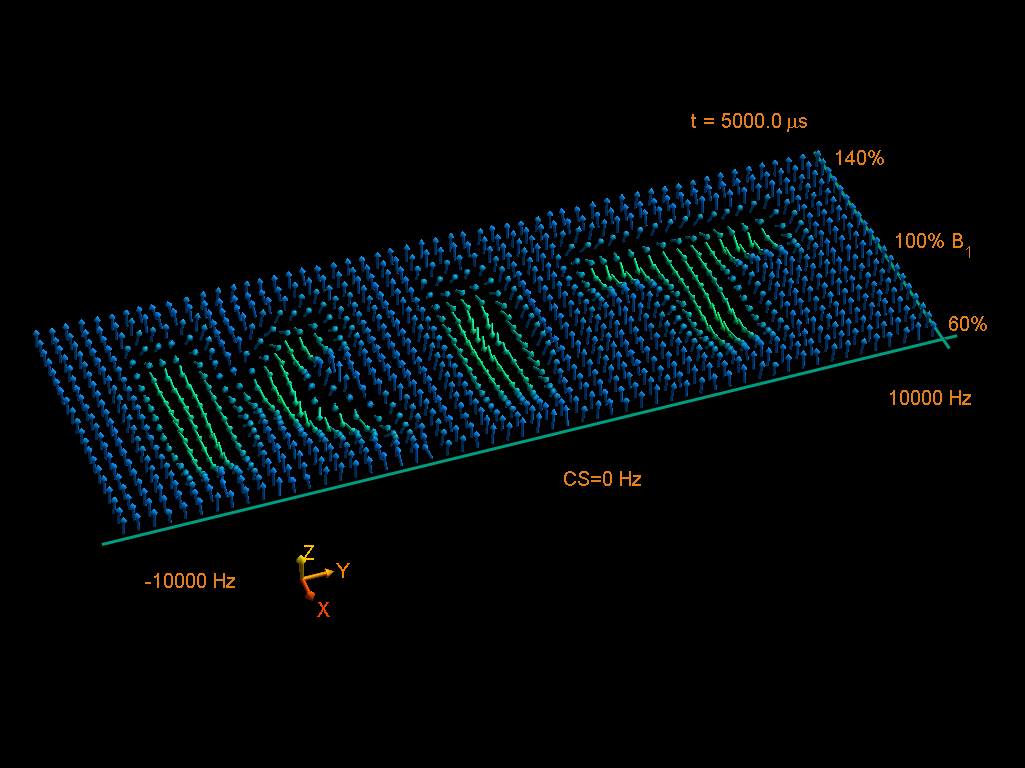
Optimal Control
Using Optimal Control Theory and the GRAPE algorithm for revolutionizing pulse trains with thousands of independent parameters.
link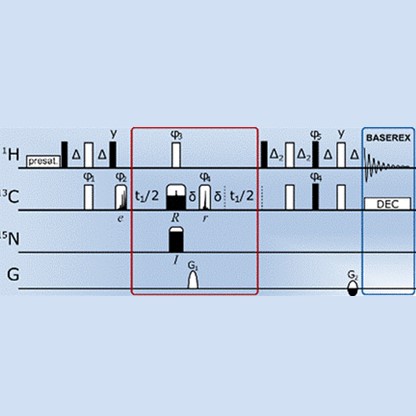
Pulse Sequences
Developing pulse sequences with utmost performance for fast and accurate determination of NMR parameters.
link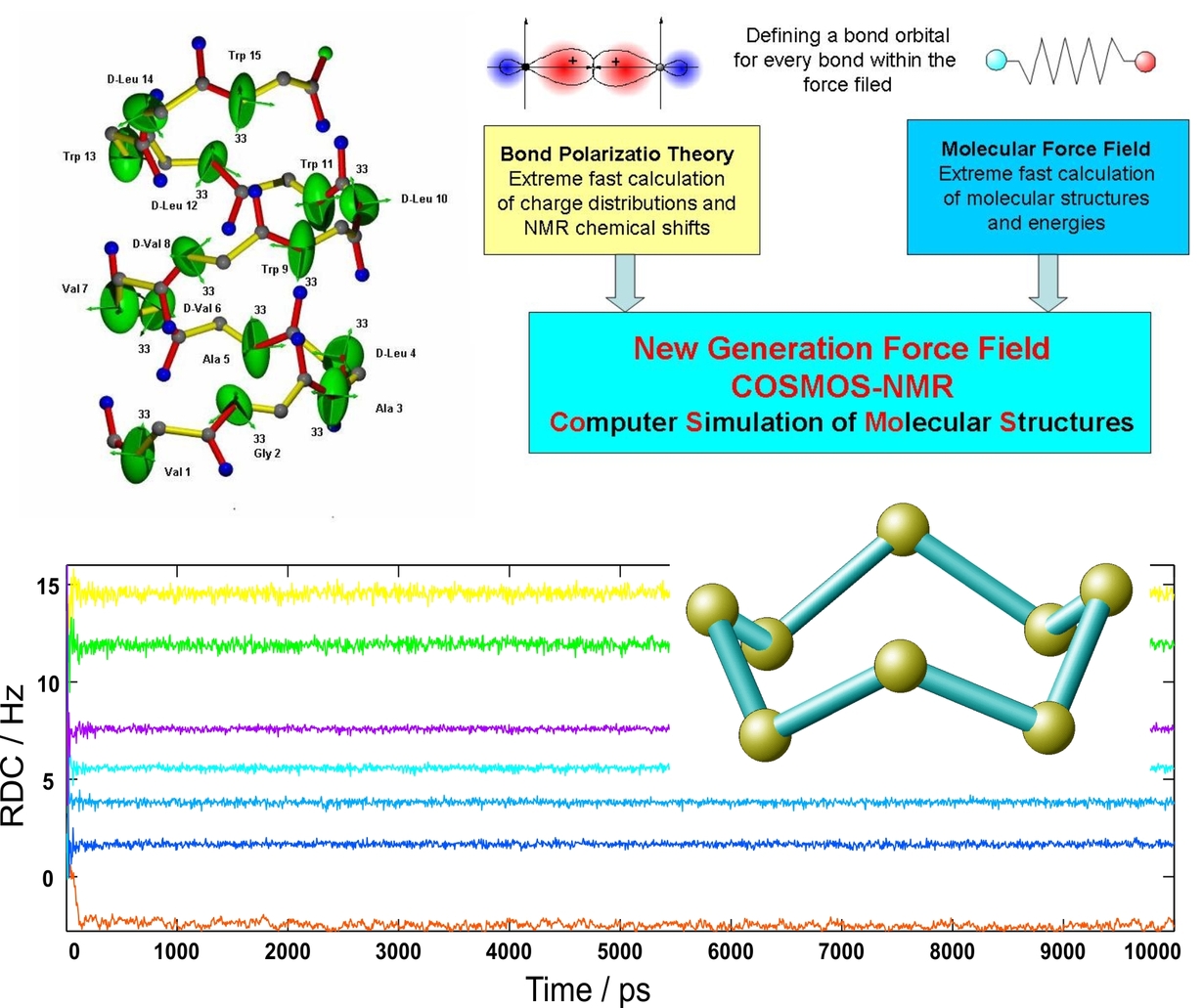
Computational Chemistry
Studying flexible molecules and chirality by newly developed molecular dynamics approaches.
link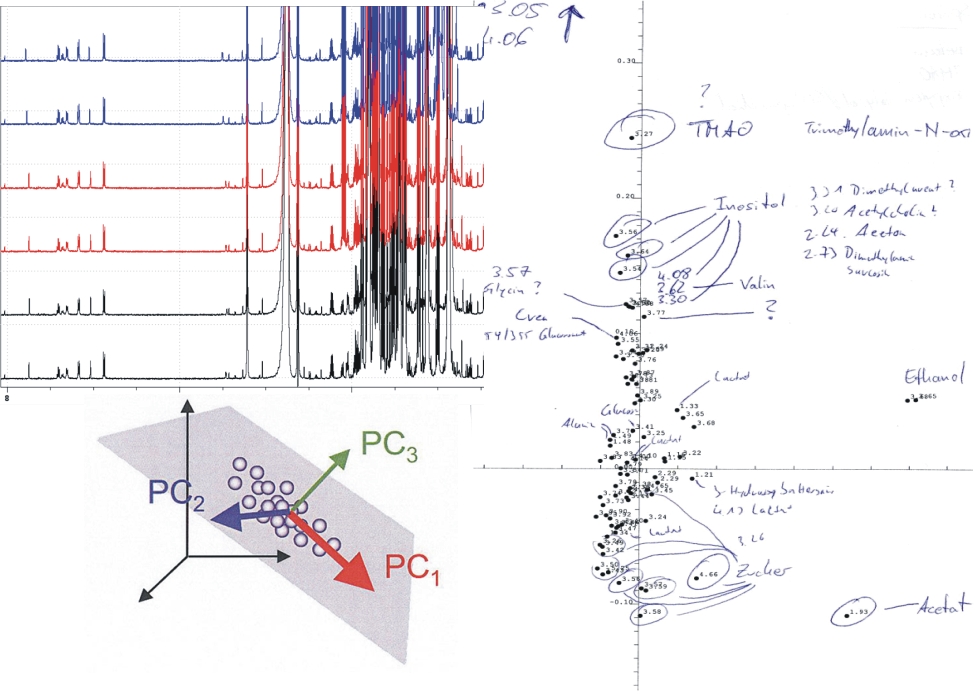
Metabolomics
Quantitatively characterizing the metabolome of organisms, organs, and biofluids by NMR spectroscopy.
link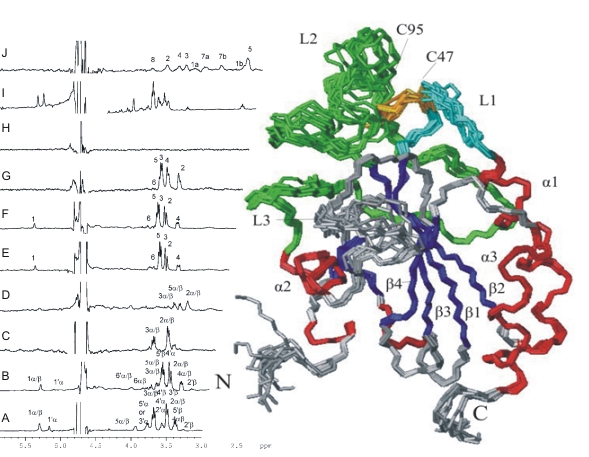
Structural Biology
Deciphering the relation of molecular structure and function of proteins with multidimensional biomolecular NMR spectroscopy.
link
Joint Research
